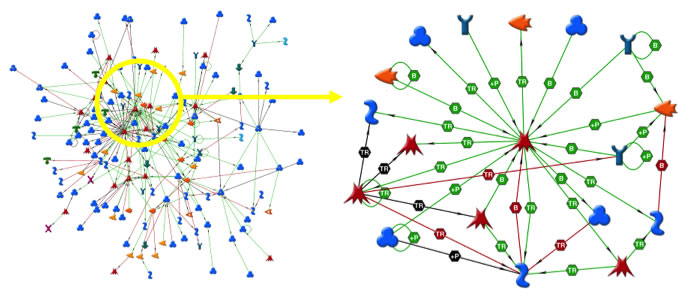
PNS versus CNS Regeneration
This project springs from the fact the axons in the peripheral nervous system regenerate well, while axons in the central nervous system regenerate poorly. First we performed subtractive hybridization experiments with cDNA libraries we made to identify genes that were differentially regulated between dorsal root ganglion (DRG) neurons and cerebellar granule neurons (CGN). We also analyzed microarray data from a similar DRG vs CGN comparison. This provided a list of about 900 unique genes for which we have 1300 unique full-length clones in expression vectors from a large collection (over 16,000 cDNAs). Of particular interest is a sub-list of 40 transcription factors (TFs) we have identified by promoter analysis and Genego Metacore network analysis as likely to regulate DRG neuron gene expression. We have tested these genes on growth permissive (laminin, polylysine) and growth inhibiting substrates (myelin and CSPGs). In a proof of principle experiment the top TF has been analyzed in detail in vitro and in vivo and we have confirmed it is differentially expressed and enhances neurite growth when overexpressed in CNS neurons. The comparison of peripheral to central nervous system neurons was continued in the projects detailed on the RNA-Seq page. The use of transcription factors to drive neuronal outgrowth in CNS neurons has led to the project detailed on the Master Nodes page.